Tutorial: Model Comparison#
This tutorial demonstrates how to compare IWFM models and simulation results using pyiwfm’s comparison tools.
Learning Objectives#
By the end of this tutorial, you will be able to:
Compare mesh differences between two models
Compare stratigraphy changes
Compute performance metrics (RMSE, NSE, etc.)
Generate comparison reports in multiple formats
Visualize differences
Setup: Create Two Model Versions#
Let’s create two versions of a model - an “original” and a “modified” version:
import numpy as np
from pyiwfm.core.mesh import AppGrid, Node, Element
from pyiwfm.core.stratigraphy import Stratigraphy
def create_base_model():
"""Create the original model."""
nodes = {}
node_id = 1
for j in range(4):
for i in range(4):
is_boundary = (i == 0 or i == 3 or j == 0 or j == 3)
nodes[node_id] = Node(
id=node_id,
x=float(i * 100),
y=float(j * 100),
is_boundary=is_boundary,
)
node_id += 1
elements = {}
elem_id = 1
for j in range(3):
for i in range(3):
n1 = j * 4 + i + 1
n2 = n1 + 1
n3 = n2 + 4
n4 = n1 + 4
elements[elem_id] = Element(
id=elem_id,
vertices=(n1, n2, n3, n4),
subregion=1,
)
elem_id += 1
grid = AppGrid(nodes=nodes, elements=elements)
grid.compute_connectivity()
# Stratigraphy
n_nodes = grid.n_nodes
strat = Stratigraphy(
n_layers=2,
n_nodes=n_nodes,
gs_elev=np.full(n_nodes, 100.0),
top_elev=np.column_stack([np.full(n_nodes, 100.0), np.full(n_nodes, 50.0)]),
bottom_elev=np.column_stack([np.full(n_nodes, 50.0), np.full(n_nodes, 0.0)]),
active_node=np.ones((n_nodes, 2), dtype=bool),
)
return grid, strat
def create_modified_model():
"""Create a modified version of the model."""
grid, strat = create_base_model()
# Modify node 6 location (interior node)
grid.nodes[6] = Node(
id=6,
x=105.0, # Moved from 100.0
y=105.0, # Moved from 100.0
is_boundary=False,
)
# Add a new node
grid.nodes[17] = Node(id=17, x=350.0, y=150.0, is_boundary=True)
# Change element 5 subregion
grid.elements[5] = Element(
id=5,
vertices=grid.elements[5].vertices,
subregion=2, # Changed from 1
)
# Modify stratigraphy
strat.gs_elev[5] = 105.0 # Raise ground surface at node 6
strat.active_node[0, 1] = False # Deactivate node 1 in layer 2
return grid, strat
# Create both versions
grid1, strat1 = create_base_model()
grid2, strat2 = create_modified_model()
print(f"Original: {grid1.n_nodes} nodes, {grid1.n_elements} elements")
print(f"Modified: {grid2.n_nodes} nodes, {grid2.n_elements} elements")
Part 1: Comparing Meshes#
Use ModelDiffer to compare the meshes:
from pyiwfm.comparison import ModelDiffer, MeshDiff, DiffType
# Create differ
differ = ModelDiffer()
# Compare meshes
mesh_diff = differ.diff_meshes(grid1, grid2)
print("Mesh Comparison Results:")
print(f" Identical: {mesh_diff.is_identical}")
print(f" Nodes added: {mesh_diff.nodes_added}")
print(f" Nodes removed: {mesh_diff.nodes_removed}")
print(f" Nodes modified: {mesh_diff.nodes_modified}")
print(f" Elements added: {mesh_diff.elements_added}")
print(f" Elements removed: {mesh_diff.elements_removed}")
print(f" Elements modified: {mesh_diff.elements_modified}")
Viewing Detailed Changes#
# List all changes
print("\nDetailed Changes:")
for item in mesh_diff.items:
print(f" {item}")
# Filter by type
added_items = [i for i in mesh_diff.items if i.diff_type == DiffType.ADDED]
modified_items = [i for i in mesh_diff.items if i.diff_type == DiffType.MODIFIED]
print(f"\nAdded items ({len(added_items)}):")
for item in added_items:
print(f" {item.path}: {item.new_value}")
print(f"\nModified items ({len(modified_items)}):")
for item in modified_items:
print(f" {item.path}: {item.old_value} -> {item.new_value}")
Part 2: Comparing Stratigraphy#
Compare stratigraphy differences:
from pyiwfm.comparison import StratigraphyDiff
# Compare stratigraphy
strat_diff = differ.diff_stratigraphy(strat1, strat2)
print("Stratigraphy Comparison Results:")
print(f" Identical: {strat_diff.is_identical}")
print(f" Number of changes: {len(strat_diff.items)}")
# List changes
print("\nStratigraphy Changes:")
for item in strat_diff.items:
print(f" {item}")
Part 3: Combined Model Diff#
Combine mesh and stratigraphy diffs into a single ModelDiff:
from pyiwfm.comparison import ModelDiff
# Create combined diff
model_diff = differ.diff(
mesh1=grid1,
mesh2=grid2,
strat1=strat1,
strat2=strat2,
)
# Get summary
print(model_diff.summary())
# Get statistics
stats = model_diff.statistics()
print(f"\nStatistics: {stats}")
Filtering Differences#
# Filter by path (e.g., only node changes)
node_changes = model_diff.filter_by_path("nodes")
print(f"Node changes: {len(node_changes.items)}")
# Filter by type (e.g., only added items)
added_only = model_diff.filter_by_type(DiffType.ADDED)
print(f"Added items: {len(added_only.items)}")
# Convert to dictionary
diff_dict = model_diff.to_dict()
print(f"Dict keys: {list(diff_dict.keys())}")
Part 4: Computing Performance Metrics#
Compare simulation results using statistical metrics:
from pyiwfm.comparison.metrics import (
ComparisonMetrics,
rmse, mae, nash_sutcliffe, percent_bias, scaled_rmse,
)
# Synthetic observed and simulated head data
rng = np.random.default_rng(42)
observed_heads = 50 + 10 * rng.standard_normal(100)
simulated_heads = observed_heads + 2 * rng.standard_normal(100)
# Calculate individual metrics
print("Individual Metrics:")
print(f" RMSE: {rmse(observed_heads, simulated_heads):.3f} ft")
print(f" Scaled RMSE: {scaled_rmse(observed_heads, simulated_heads):.4f}")
print(f" MAE: {mae(observed_heads, simulated_heads):.3f} ft")
print(f" NSE: {nash_sutcliffe(observed_heads, simulated_heads):.3f}")
print(f" PBIAS: {percent_bias(observed_heads, simulated_heads):.2f}%")
Computing All Metrics#
# Compute all metrics at once
metrics = ComparisonMetrics.compute(observed_heads, simulated_heads)
# Print summary
print(metrics.summary())
# Get rating
print(f"\nModel Rating: {metrics.rating()}")
# Convert to dictionary
metrics_dict = metrics.to_dict()
print(f"\nAs dict: {metrics_dict}")
Part 5: Time Series Comparison#
Compare time series data:
from pyiwfm.comparison.metrics import TimeSeriesComparison
# Create example time series
times = np.arange(365) # Daily data for one year
observed = 50 + 10 * np.sin(times * 2 * np.pi / 365) # Seasonal pattern
simulated = observed + 2 * np.random.randn(365) # Add noise
# Create comparison
ts_comparison = TimeSeriesComparison(
times=times,
observed=observed,
simulated=simulated,
)
print(f"Time Series Comparison:")
print(f" Points: {ts_comparison.n_points}")
print(f" Valid points: {ts_comparison.n_valid_points}")
print(f" RMSE: {ts_comparison.metrics.rmse:.3f}")
print(f" NSE: {ts_comparison.metrics.nash_sutcliffe:.3f}")
# Get residuals
residuals = ts_comparison.residuals
print(f" Mean residual: {residuals.mean():.3f}")
print(f" Std residual: {residuals.std():.3f}")
Part 6: Spatial Comparison#
Compare spatial fields:
from pyiwfm.comparison.metrics import SpatialComparison
# Coordinates
x = np.array([grid1.nodes[i].x for i in sorted(grid1.nodes.keys())])
y = np.array([grid1.nodes[i].y for i in sorted(grid1.nodes.keys())])
# Head values from two simulations
heads_run1 = 50 + 0.01 * x + 0.02 * y
heads_run2 = heads_run1 + np.random.randn(len(x))
# Create spatial comparison
spatial = SpatialComparison(
x=x, y=y,
observed=heads_run1,
simulated=heads_run2,
)
print(f"Spatial Comparison:")
print(f" Points: {spatial.n_points}")
print(f" RMSE: {spatial.metrics.rmse:.3f}")
# Get error field
errors = spatial.error_field
print(f" Max error: {np.max(np.abs(errors)):.3f}")
print(f" Mean error: {np.mean(errors):.3f}")
# Relative errors
rel_errors = spatial.relative_error_field
print(f" Max relative error: {np.max(np.abs(rel_errors)) * 100:.1f}%")
Regional Analysis#
# Define regions (e.g., by subregion)
regions = np.array([
grid1.elements[e].subregion
for e in sorted(grid1.elements.keys())
for _ in range(len(grid1.elements[e].vertices))
])[:len(x)] # Match array size
# In practice, assign region to each node
regions = np.ones(len(x), dtype=int)
regions[x > 150] = 2 # Simple split by x coordinate
# Compute metrics by region
regional_metrics = spatial.metrics_by_region(regions)
print("\nMetrics by Region:")
for region_id, region_metrics in regional_metrics.items():
print(f" Region {region_id}:")
print(f" RMSE: {region_metrics.rmse:.3f}")
print(f" NSE: {region_metrics.nash_sutcliffe:.3f}")
Part 7: Generating Reports#
Generate comparison reports in various formats:
from pyiwfm.comparison.report import (
ReportGenerator,
TextReport,
JsonReport,
HtmlReport,
ComparisonReport,
)
Text Report#
# Create text report
text_report = TextReport()
content = text_report.generate(model_diff)
print(content[:500]) # Print first 500 chars
# Save to file
text_report.save(model_diff, "comparison_report.txt")
JSON Report#
import json
# Create JSON report
json_report = JsonReport(indent=2)
content = json_report.generate(model_diff)
# Parse to verify
data = json.loads(content)
print(f"JSON keys: {list(data.keys())}")
# Save to file
json_report.save(model_diff, "comparison_report.json")
HTML Report#
# Create HTML report
html_report = HtmlReport(title="Model Comparison Report")
content = html_report.generate(model_diff)
# Save to file
html_report.save(model_diff, "comparison_report.html")
print("Saved comparison_report.html")
Using ReportGenerator#
The ReportGenerator provides a unified interface:
# Create report generator
generator = ReportGenerator()
# Generate in different formats
text_content = generator.generate(model_diff, format="text")
json_content = generator.generate(model_diff, format="json")
html_content = generator.generate(model_diff, format="html")
# Auto-detect format from file extension
generator.save(model_diff, "report.txt") # Text format
generator.save(model_diff, "report.json") # JSON format
generator.save(model_diff, "report.html") # HTML format
Comprehensive Report#
Create a report with both model diffs and metrics:
# Create comprehensive report
report = ComparisonReport(
title="Model Version Comparison",
description="Comparing original model to modified version",
model_diff=model_diff,
head_metrics=metrics,
)
# Generate outputs
print(report.to_text())
# Save in different formats
report.save("full_report.txt")
report.save("full_report.json")
report.save("full_report.html")
Part 8: Visualizing Differences#
Visualize the comparison results:
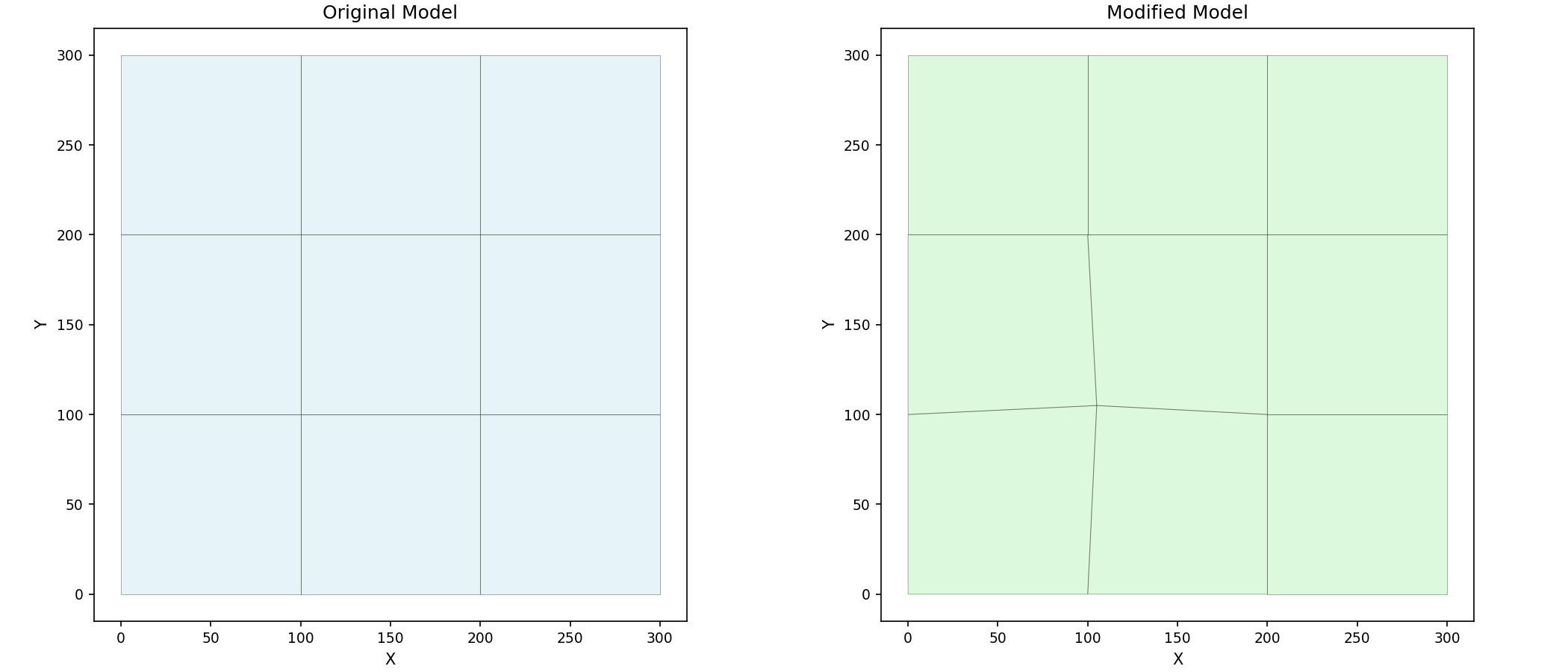
# Create side-by-side comparison plot
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
# Original model
plot_mesh(grid1, ax=axes[0], show_edges=True, fill_color="lightblue")
axes[0].set_title("Original Model")
# Modified model
plot_mesh(grid2, ax=axes[1], show_edges=True, fill_color="lightgreen")
axes[1].set_title("Modified Model")
plt.show()
Plotting Residuals#
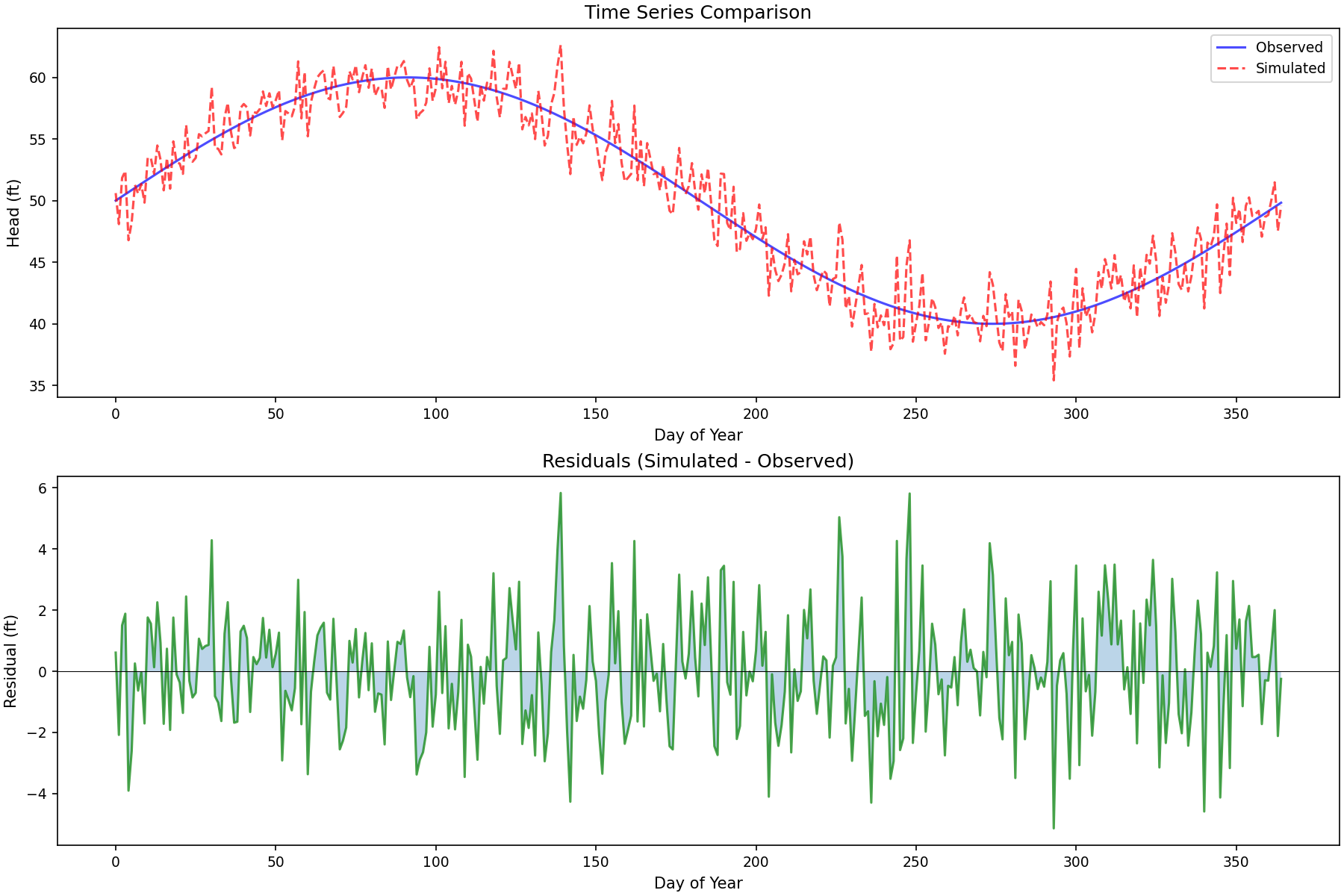
# Time series residual plot
fig, axes = plt.subplots(2, 1, figsize=(12, 8))
# Time series plot
axes[0].plot(times, observed, 'b-', label='Observed', alpha=0.7)
axes[0].plot(times, simulated, 'r--', label='Simulated', alpha=0.7)
axes[0].set_xlabel('Day of Year')
axes[0].set_ylabel('Head (ft)')
axes[0].legend()
axes[0].set_title('Time Series Comparison')
# Residuals plot
axes[1].plot(times, residuals, 'g-', alpha=0.7)
axes[1].axhline(y=0, color='k', linestyle='-', linewidth=0.5)
axes[1].fill_between(times, residuals, 0, alpha=0.3)
axes[1].set_xlabel('Day of Year')
axes[1].set_ylabel('Residual (ft)')
axes[1].set_title('Residuals (Simulated - Observed)')
plt.show()
Summary#
This tutorial covered:
Mesh comparison: Detect node/element changes
Stratigraphy comparison: Detect layer changes
Performance metrics: RMSE, Scaled RMSE, MAE, NSE, PBIAS
Time series: Compare temporal data with residual analysis
Spatial comparison: Regional analysis of spatial fields
Report generation: Text, JSON, and HTML reports
Key classes:
ModelDiffer- Compare modelsMeshDiff,StratigraphyDiff- Diff containersComparisonMetrics- Statistical metricsReportGenerator- Multi-format reports